Lecture 5 Video 4
🧬 Ligand–Protein Interactions by NMR – Lecture 5 Video 4 Summary
This lecture continues the discussion of how NMR can be used to study ligand–protein interactions, moving from chemical shift perturbations to the more structurally powerful method of NOEs (Nuclear Overhauser Effects) .
Below is a structured and detailed walkthrough of all key concepts.
1️⃣ Chemical Shift Perturbation vs. NOEs
🔹 Chemical Shift Perturbation (CSP)
Previously, you learned that ligand binding often causes chemical shift changes in protein resonances.
What CSP tells you:
- Which residues are affected by binding
- Which region of the protein is involved
- Rough binding site location
What CSP does NOT give you:
- Exact atomic distances
- 3D structure of the complex
CSP is excellent for mapping the interface, but it cannot define the detailed structure of the bound complex .
🔹 NOEs (Nuclear Overhauser Effects)
This is where things become structurally powerful.
NOEs provide:
- Direct distance restraints
- Information about atoms that are close in space (~<5 Å)
- The possibility to calculate a full 3D structure of the complex
But there is a catch.
⚠️ Requirement for NOEs
To observe NOEs between ligand and protein:
- The binding must be strong
- The complex must have a long lifetime
- Exchange must not be too fast
If binding is weak or transient, intermolecular NOEs are too weak or averaged out .
2️⃣ Example: Fatty Acid Binding Protein (FABP)
The lecture presents a real research example involving the human brain-type fatty acid binding protein (FABP) .
🧱 Structure of FABP

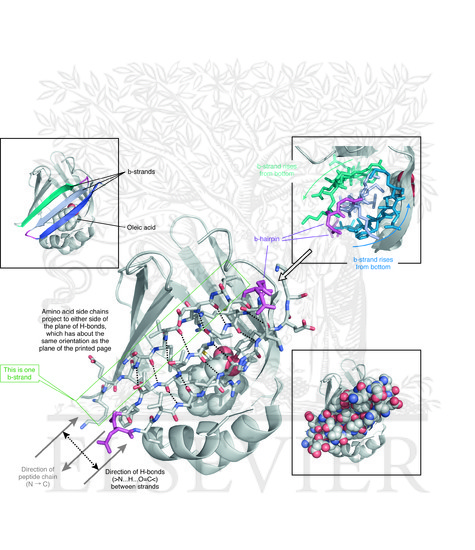

Structural features:
- A β-barrel protein
- A “lid” composed of two α-helices
- Internal hydrophobic cavity
This architecture is ideal for binding hydrophobic molecules.
3️⃣ Why Does It Bind Fatty Acids Inside?
Fatty acids are:
- Long
- Hydrophobic
- Poorly soluble in water
The protein’s role:
- Bind fatty acids
- Transport them in aqueous environments
💡 Key principle:
Hydrophobic molecules are “hidden” inside the protein to prevent exposure to water.
The β-barrel acts as:
- A protective pocket
- A transport container
This is a beautiful example of structure-function relationship.
4️⃣ Intermolecular NOEs in This Study
Researchers added fatty acids and measured intermolecular NOEs.
They observed:
- NOEs between fatty acid atoms and protein atoms
- All interacting atoms were located inside the barrel
These atoms were visualized as black spheres in the structure .
🔬 What does this prove?
- The fatty acid binds inside the barrel
- Not on the surface
- Not in multiple sites
This is direct spatial evidence.
5️⃣ Structural Calculation from NOEs
Because sufficient NOEs were observed, they could:
- Use NOE-derived distance restraints
- Calculate a structure of the complex
- Generate a bundle of structures
Why a bundle?
NMR structures are typically represented as:
- ~20 lowest-energy conformers
- All consistent with experimental restraints
The lecture notes that:
- The fatty acid is not super well defined
- But the binding location is clear
This is typical for small ligands — they may retain some flexibility.
6️⃣ Specific Binding Interactions
The structure revealed specific interactions:
🔹 Carboxylate group of fatty acid:
- Close to an arginine side chain
- Close to a tyrosine
This implies:
- Charge–charge interaction (carboxylate ↔ arginine)
- Hydrogen bonding (carboxylate ↔ tyrosine)
These interactions:
- Stabilize binding
- Anchor the ligand in a defined orientation
Importantly: These interactions were known from previous studies, and the NMR data confirmed them .
This is an example of experimental validation.
7️⃣ Structural Interpretation: What We Learn
From this example, you should understand several major concepts:
🧠 Concept 1: Binding Requires Complementarity
- Hydrophobic ligand
- Hydrophobic cavity
- Polar head group
- Charged residues inside cavity
This is chemical complementarity.
🧠 Concept 2: NOEs Provide Spatial Information
Unlike chemical shifts:
- NOEs give distance restraints
- Allow structure calculation
- Define binding orientation
That’s the key upgrade.
🧠 Concept 3: Strong Binding is Necessary
NOEs require:
- Long-lived complex
- Slow/intermediate exchange
- Sufficient population of bound state
Weak binders → CSP mapping only Strong binders → possible full structure
🧠 Concept 4: NMR Structures Are Ensembles
The lecture shows:
- A bundle of 20 fatty acid conformations
- Some structural variability
- But consistent binding pocket
NMR does not give a single structure. It gives an ensemble consistent with experimental restraints.
8️⃣ Big Picture Summary
This lecture teaches an important progression:
| Method | What You Learn | Structural Detail |
|---|---|---|
| Chemical shift perturbation | Binding site region | Low resolution |
| Intermolecular NOEs | Exact contacts | High resolution |
| Structure calculation | Full complex structure | Atomic detail |
🎯 Key Takeaways
- Chemical shifts map binding sites but don’t give structures.
- NOEs require strong binding.
- Intermolecular NOEs give distance restraints.
- Strong NOE data enables structure calculation.
- FABP binds fatty acids inside its β-barrel.
- The fatty acid is stabilized by electrostatic and hydrogen bonding interactions.
- NMR structures are ensembles, not single models.