Lecture 1 Video 6 Optical Activity Summary
🌈 Polarized Light & Chirality: The Big Picture
Light can be polarized, meaning its electric field oscillates in a defined way. Two important forms here are:
- Linearly polarized light
- Circularly polarized light (left-handed vs right-handed)
Most molecules interact identically with left- and right-handed circularly polarized light. 👉 Chiral (asymmetric) molecules do not — and that asymmetry is the foundation of both optical rotation and circular dichroism.
🔄 Optical Rotation (OR)
What is it?
When linearly polarized light passes through a chiral medium (e.g. sugar solution), the plane of polarization rotates.
- The rotation angle is usually called α
- It depends on:
- Path length (ℓ)
- Wavelength (λ)
- Difference in refractive indices for left vs right circularly polarized light
Why does this happen?
Linearly polarized light can be seen as a sum of left- and right-handed circularly polarized light. If these two components travel at different speeds, the polarization plane rotates.
📈 Optical Rotary Dispersion (ORD)
If you plot optical rotation vs wavelength, you get an ORD spectrum.
- Often denoted φ(λ)
- Shows how optical rotation changes with wavelength
- Useful, but limited for proteins
🌀 Circular Dichroism (CD): The Star of the Show
Core idea
Circular dichroism measures the difference in absorption between:
- Left-handed circularly polarized light
- Right-handed circularly polarized light
This only happens if the molecule is chiral.
🧪 Absorbance refresher (Beer–Lambert law)
A = logleft( rac{I_0}{I} ight) = arepsilon cdot c cdot ell
In CD:
- Left and right circularly polarized light have different absorbances
- This means different extinction coefficients
- The measurable quantity is: Delta A = A_L - A_R
🟠 Why Ellipticity? (CD Units Explained)
Instead of reporting ΔA directly, CD uses ellipticity (θ).
Physical interpretation
- If left and right components are absorbed equally → light stays linear → θ = 0
- If absorption differs → resultant electric field traces an ellipse
- The angle θ describes how “elliptical” the light becomes
Mathematical relationship
For small angles (almost always true):
heta ( ext{degrees}) = 32.98 cdot Delta A
✔️ Ellipticity is directly proportional to the absorbance difference
📏 Mean Residue Ellipticity (Protein CD Units 😵💫)
Spectroscopists normalize CD data to make proteins comparable.
They define mean residue ellipticity, which accounts for:
- Observed ellipticity
- Protein concentration
- Path length
- Molecular weight
- Number of amino acid residues
Resulting unit:
ext{deg·cm}^2· ext{dmol}^{-1}· ext{residue}^{-1}
Weird unit — but standard in protein CD literature.
🧬 Why CD Is So Powerful for Proteins
UV absorption sources in proteins
- Aromatic amino acids → near UV
- Peptide bond → far UV (≈ 190–220 nm)
Proteins are chiral polymers (except glycine), so they show strong CD signals in the far-UV region.
🧠 Secondary Structure Signatures in CD
Different secondary structures give distinct CD spectra:
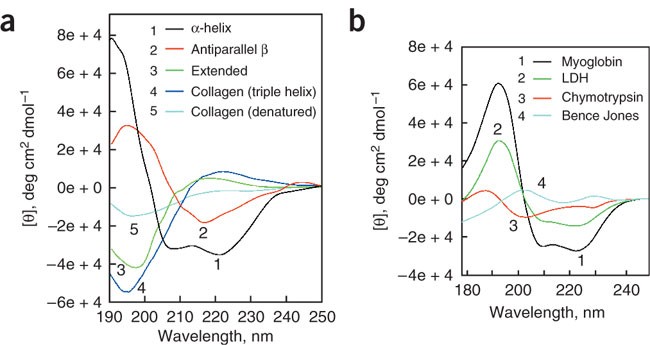


α-Helix 🌀
- Negative bands: 208 nm & 222 nm
- Strong positive band: ~185 nm
β-Sheet 🧵
- Bands shifted relative to helices
- Different positive/negative pattern
Random coil 🎲
- Completely different shape
➕ Spectra Are Additive!
A protein with:
- 50% β-sheet
- 30% α-helix
- 20% random coil
will show a weighted sum of those three spectra.
👉 This allows quantitative estimation of secondary structure content.
🔥 Following Protein Folding & Unfolding
CD can monitor:
- Thermal unfolding
- Chemical denaturation
- Ligand binding
- Structural stability
You simply track how the CD signal changes with:
- Temperature
- Time
- Additives
🧪 Real Protein Examples



- Myoglobin → almost purely α-helical → CD matches helix signature
- Triose phosphate isomerase → mainly β-sheet but mixed → intermediate spectrum
- Mixed α/β proteins → composite spectra
⚠️ Practical Limitations & Sample Preparation
CD is powerful but experimentally demanding, especially at low wavelengths.
Key problems
- Buffers may absorb UV light
- Even if buffer CD = 0, strong absorption kills signal quality
- You’re trying to detect tiny differences in a small remaining signal
Best practices ✅
- Use very pure protein
- Minimize buffer concentration
- Avoid:
- Metal ions
- Halides (especially Br⁻, I⁻)
- With care, measurements down to ~190 nm are achievable
🧠 Final Take-Home Messages
- Optical rotation → refractive index differences
- Circular dichroism → absorption differences
- Ellipticity quantifies CD
- Far-UV CD is a gold standard for:
- Secondary structure analysis
- Protein folding studies
- CD spectra are additive, enabling structural estimation
- Sample preparation is critical