Lecture 1 Video 5 Protein Fluorescence Summary
🌈 Protein Fluorescence: Principles & Applications
Protein fluorescence is a powerful, sensitive spectroscopic tool used to study protein structure, folding, and stability. The key idea is simple: some molecules absorb light and re-emit it at a longer wavelength, and proteins conveniently contain amino acids that can do this.
🔦 1. How Is Fluorescence Measured?
Basic optical setup
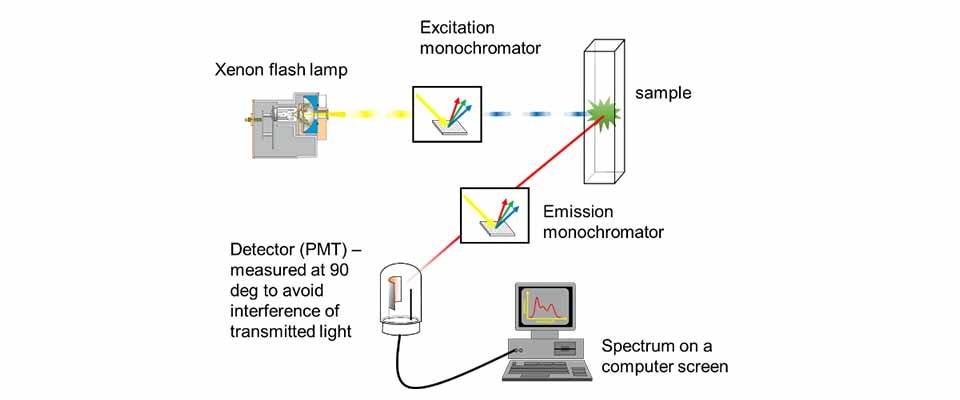
A typical fluorescence experiment is based on optical spectroscopy and consists of:
- Light source (lamp) – provides excitation light
- Monochromator (excitation) – selects a specific wavelength
- Sample – absorbs some light
- Detection system – measures either transmitted or emitted light
Absorption vs fluorescence geometry
- Absorption spectroscopy measures light that passes straight through the sample.
- Fluorescence spectroscopy measures light emitted at a 90° angle relative to the incoming beam.
🔑 Why 90°? To minimize the amount of direct lamp light reaching the detector. This greatly improves the signal-to-noise ratio, since fluorescence is much weaker than excitation light.

🔄 2. Emission Scans vs Excitation Scans
There are two main ways to record fluorescence data:
📈 Emission scan (most common for proteins)
- Keep excitation wavelength fixed
- Scan emitted wavelengths
- Result: spectrum of emitted light for a given excitation
👉 This is what you most often see in protein fluorescence literature.
📉 Excitation scan
- Keep emission wavelength fixed
- Scan excitation wavelengths
- Result: tells you which excitation wavelengths lead to fluorescence
✨ 3. Fluorophores: The Source of Fluorescence
A fluorophore is any chemical group that fluoresces.
🧬 Intrinsic fluorophores (naturally present in proteins)
Proteins contain three aromatic amino acids that can fluoresce:
| Amino acid | Fluorescence strength | Notes |
|---|---|---|
| Phenylalanine (Phe) | Very weak ❌ | Hard to measure |
| Tyrosine (Tyr) | Moderate ⚠️ | Only useful if no Trp |
| Tryptophan (Trp) | Very strong ✅ | Dominates signal |
🔑 Key rule: If tryptophan is present, its fluorescence overwhelms tyrosine and phenylalanine.
🧪 Extrinsic fluorophores
If a protein does not contain suitable intrinsic fluorophores, you can:
- Chemically attach a fluorescent probe
- Introduce fluorescence artificially
These are called extrinsic fluorophores, and they are widely used because fluorescence is extremely sensitive.
🌊 4. Why Tryptophan Is So Powerful
Tryptophan is the gold standard of intrinsic protein fluorescence because:
- It absorbs light strongly
- It fluoresces efficiently
- Its emission wavelength is environment-dependent
🔬 Sensitivity to solvent polarity
The emission maximum of tryptophan shifts depending on its surroundings:
- Polar environment (water) → higher wavelength
- Apolar environment (protein core) → lower wavelength

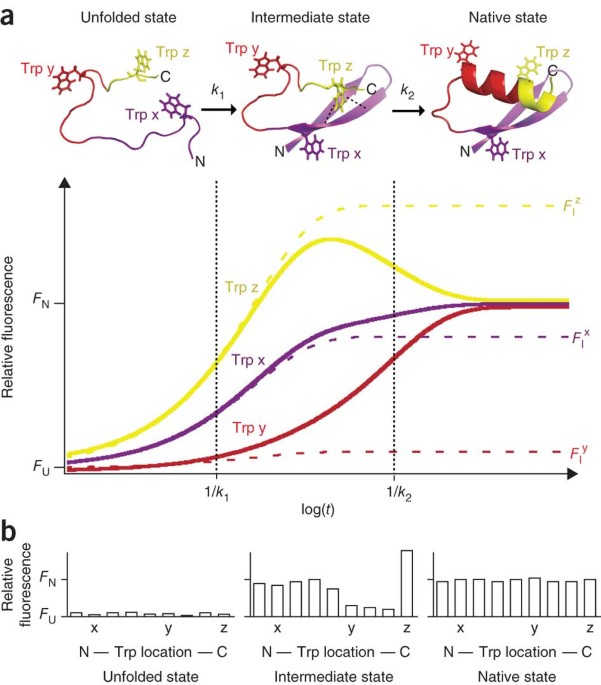
🧩 5. Protein Folding and Unfolding
This environmental sensitivity makes tryptophan ideal for studying protein folding.
Folded protein
- Tryptophan is usually buried in the hydrophobic core
- Emits at ~325–335 nm
Unfolded protein
- Tryptophan becomes exposed to water
- Emits at ~350–355 nm
📌 Key insight: Protein unfolding causes a red shift (increase in emission wavelength).
📊 6. Following Protein Stability with Fluorescence
Because fluorescence is highly sensitive, you can monitor folding transitions using very small amounts of protein.
Typical experiment
- Measure Trp fluorescence at a fixed wavelength (e.g. 355 nm)
- Change an external condition:
- 🌡 Temperature
- ⚗ pH
- 🧪 Denaturant concentration (urea, guanidinium chloride)
- Plot fluorescence intensity vs condition
Result
You obtain a sigmoidal unfolding curve:
- Folded baseline
- Unfolding transition
- Unfolded baseline
The midpoint gives:
- Melting temperature (Tₘ) for thermal unfolding
- Midpoint pH or denaturant concentration for chemical unfolding
🎯 This allows you to quantify protein stability in solution.
🧠 Big Picture Takeaways
- Fluorescence is measured at 90° to reduce background light
- Emission scans are most common in protein studies
- Proteins have intrinsic fluorophores, especially tryptophan
- Tryptophan emission wavelength reports on local environment
- Protein unfolding causes a shift from ~330 nm → ~350 nm
- Fluorescence enables high-sensitivity monitoring of folding, stability, and denaturation