Day 7 part 3
🧪 Fun & Educational Summary: Gel Chromatography / Size-Exclusion Chromatography
This technique is one of the most elegant protein purification methods because it separates proteins based purely on size in solution, without relying on charge or affinity.
It is often called:
- Gel filtration chromatography
- Size-exclusion chromatography (SEC)
- Molecular sieve chromatography
All mean essentially the same thing.
🎯 Core Principle of Gel Chromatography
Yes — your understanding is correct:
larger proteins come out first
This is the key principle.
The separation happens because the column contains porous beads.
Think of the beads like tiny sponges filled with tunnels and pores.



When proteins flow through the column:
- large proteins cannot enter many pores
- medium proteins enter some pores
- small proteins enter many pores
Because of this:
- large proteins take a shorter path
- small proteins take a longer path
So:
large proteins elute firstsmall proteins elute last
This is exactly what the file describes.
🚶 Why Do Larger Proteins Come Out First?
This is the most important conceptual point.
Imagine two people moving through a city:
- one takes the main road
- one keeps entering side streets
Who reaches the destination first?
The one on the main road.
That is exactly SEC.
Large proteins stay mostly in the space between beads.
Small proteins diffuse in and out of pores many times.
This delays them.
So separation is based on:
how much time molecules spend inside bead pores
🧬 Is Separation Based on Molecular Weight or Hydrodynamic Size?
Excellent question.
The most correct answer is:
hydrodynamic size
This is more precise than molecular weight.
The file mentions both, but the physically correct parameter is:
hydrodynamic radius / Stokes radius
Why not just molecular weight?
Because proteins with the same molecular weight can have different shapes.
Example:
- a compact globular protein
- an elongated fibrous protein
Even if both are 100 kDa, the elongated one behaves as larger in solution.
So SEC actually separates based on:
effective size in solution
This includes:
- molecular mass
- shape
- hydration shell
That is what hydrodynamic size means.
This distinction is very important at master’s level.
🧫 What Are the Beads Made Of?
You asked whether the matrix is made of dextran and cross-linked agarose.
Yes — commonly, yes.
The file mentions several possible matrices:
- dextran
- agarose
- polyacrylamide
- combinations / cross-linked hybrids
Common commercial examples:
- Sephadex → cross-linked dextran
- Sepharose → agarose-based
- Superdex → dextran + agarose hybrid
Why these materials?
Because they need to be:
- water-insoluble
- chemically inert
- biocompatible
- porous
- mechanically stable
The file explicitly mentions this.
Proteins are purified in aqueous buffer, so the matrix must not dissolve.
🔗 Why Does Cross-Linking Give Rigidity?
Excellent question.
Yes — cross-linking increases rigidity.
This is a very important polymer chemistry concept.
Cross-linking means polymer chains are chemically connected to each other.
Without cross-links:
- chains move freely
- bead becomes soft / gel-like
- may collapse under pressure
With cross-links:
- chains are tied together into a network
- movement becomes restricted
- bead becomes mechanically stronger

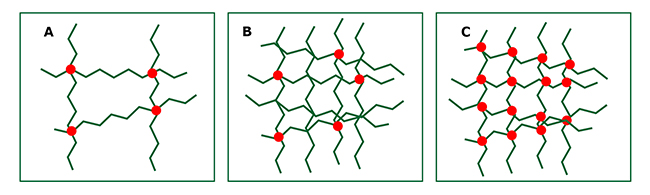

Think of it like:
- loose spaghetti = flexible
- spaghetti glued together = rigid
That is why the file says cross-links provide rigidity.
Why is rigidity important?
Because liquid is pushed through the column under pressure.
If beads collapse:
- pore size changes
- separation quality worsens
- flow rate becomes inconsistent
So rigidity ensures:
stable pore size and reproducible separation
🧪 Important Theory Beyond Your Questions
Let’s cover the terms from the file carefully.
📦 Total Volume (Vt)
The file calls this total gel volume.
V_t
This is:
entire liquid volume inside the column
It includes:
- liquid outside beads
- liquid inside pores
V_t = V_o + V_i
🌊 Void Volume (Vo)
This is one of the most important concepts.
You asked:
“Vo = volume outside the beads?”
Yes — exactly.
V_o
is the volume of mobile phase outside the pores
This means:
- between beads
- around beads
- spaces molecules travel without entering pores
This is often called:
void volumeexcluded volume
🚪 What Does Elution at Void Volume Mean?
This is a critical concept.
If a protein is too large to enter any pore, it travels only through (V_o).
That means it comes out as early as physically possible.
So:
elution at Vo means the molecule was completely excluded from the beads
This is exactly what the file says.
Important correction to your understanding
You asked:
“something about elution at the void volume?”
The precise meaning is:
the molecule is larger than the fractionation range of the column
So if both proteins are too large:
- 300 kDa
- 500 kDa
they both elute at (V_o)
and cannot be separated
This is a very common exam question.
🚫 Why Very Large Proteins Cannot Be Separated
Because both are excluded from pores.
They both travel identical paths.
So the column sees them as the same “size class”.
This is why choosing the correct column pore range is essential.
The file mentions this explicitly.
🧪 Internal Volume (Vi)
V_i
This is:
volume inside the pores of the beads
Only molecules that can enter pores access this space.
Small molecules explore more of (V_i)
Large molecules explore less.
⏱️ Elution Volume (Ve)
V_e
This is the volume of buffer passed through when the molecule elutes.
Very important relationship:
V_e = V_o + K_dV_i
The file calls this distribution coefficient / dispersion constant.
Important correction:
This (K_d) is NOT the binding dissociation constant
It is a chromatography coefficient.
That distinction is crucial.
📊 Retention Parameters in Gel Chromatography
You specifically asked about retention parameters.
The major ones are:
1) Elution volume
V_e
Measured experimentally from chromatogram peak position.
2) Void volume
V_o
Usually measured with a very large standard molecule like Blue Dextran 2000
because it cannot enter pores.
3) Distribution coefficient
K_ = rac{V_e - V_o}{V_t - V_o}
This is extremely important.
It tells how much of pore volume the molecule can access.
Range:
0 le K_ le 1
Where:
- 0 = fully excluded
- 1 = fully included
🧠 Key Conceptual Takeaway
The entire theory can be summarized as:
proteins are separated by how much pore volume they can access
Large proteins:
K_approx 0
Small proteins:
K_approx 1
🎓 Common Misunderstanding to Correct
A subtle correction:
Many students think proteins are “filtered” like a sieve membrane.
That is not quite correct.
This is not filtration by blocking.
Instead it is:
partitioning between outside and inside pore volume
That distinction matters.
The protein is not trapped.
It simply spends more or less time diffusing into pores.