Day 1 part 2
Protein chemistry
1. Genetic Code & Amino-Acid Similarity 🧬
- The genetic code consists of 64 codons:
- 61 encode amino acids
- 3 are stop codons
- Amino acids with similar chemical properties cluster together in the codon table:
- Non-polar amino acids cluster together
- Polar, basic, and acidic amino acids also cluster
Why this matters
- Single-nucleotide mutations often change an amino acid into another with similar chemistry
- This buffers proteins against catastrophic functional loss
- Example: a hydrophobic amino acid often mutates into another hydrophobic one
2. Canonical, Rare, and Non-Natural Amino Acids 🧪
- 20 standard amino acids are used in proteins
- Two rare natural amino acids:
- Selenocysteine
- Pyrrolysine
- These are not expected to be memorized, but they demonstrate that biology can expand the genetic code
Non-natural amino acids
- Can be engineered into proteins
- Retain amino + carboxyl groups but have novel side chains
- Require genetic engineering
- Used to introduce:
- Fluorescent probes
- Crosslinkers
- Novel chemical reactivity
3. Naming Systems & Mass Spectrometry ⚖️
- Amino acids have:
- Full name
- 3-letter code
- 1-letter code
- Ambiguous notation:
- Asx = Asparagine or Aspartate
- Glx = Glutamine or Glutamate
Mass spectrometry problem
- Leucine and isoleucine have:
- Identical molecular weight
- Indistinguishable by MS alone
- Result: ambiguity in protein sequencing unless supported by DNA data
4. Hydrophobicity & Transfer Free Energy 🌊


Hydrophobicity is quantified by:
ΔG of transferring an amino-acid side chain from a hydrophobic to a hydrophilic environment
Key points
- Absolute ΔG values depend on experimental setup
- Relative order is conserved
Hydrophobic extremes
- Very hydrophobic: Phenylalanine, Leucine, Isoleucine
- Borderline: Glycine (tiny side chain)
- Strongly hydrophilic: Charged amino acids
Does hydrophobicity depend on pH?
- Generally no
- Exception: amino acids with titratable side chains (e.g. Lys, Asp, Glu)
- Protonation state changes → charge changes → hydrophobicity changes
5. Ionization, pKa, and pI 🔋

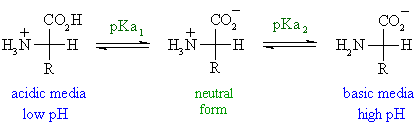
Example: Glycine
- Two ionizable groups:
- α-carboxyl (~pKa ≈ 2)
- α-amino (~pKa ≈ 9.6)
- pI ≈ 6 (net charge = 0)
Key definitions
- pKa: pH where 50% protonated
- pI: pH where net charge = 0
Amino acids with side-chain pKa
- Basic: Lys, Arg, His
- Acidic: Asp, Glu
- Special: Cys (~8.3), Tyr (~10.9)
6. Why Cysteine Is Exceptionally Reactive ⚡
This is a core concept.
pKa of cysteine ≈ 8.3
- At physiological pH (~7):
- ~10% exists as thiolate (S⁻)
Consequences
- Thiolate is a very strong nucleophile
- Protonated thiol (–SH) is weak
- Even partial deprotonation is enough for reactivity
Functional impact
- Cysteine participates in:
- Catalysis
- Redox chemistry
- Disulfide bond formation
Why tyrosine does not behave similarly
- pKa ≈ 10.9
- Almost no deprotonation at pH 7
- Negligible nucleophilicity in biology
7. Why Serine (or Cysteine) Is Placed Near Histidine 🧠
This explains catalytic triads.


- Serine and cysteine alone are weak nucleophiles
- Histidine acts as a general base
- Histidine:
- Accepts a proton
- Activates Ser-O⁻ or Cys-S⁻
Result
- Formation of a powerful nucleophile
- Enables peptide bond cleavage
- Core principle of:
- Serine proteases
- Cysteine proteases
8. pKa Is NOT Fixed in Proteins 🌡️
- pKa values listed in tables are for free amino acids in water
- In proteins, pKa shifts due to:
- Nearby charges
- Hydrophobic environments
- Hydrogen bonding
Examples
- Acidic residue near negative charges → higher pKa → prefers protonation
- Acidic residue near positive charges → lower pKa → prefers deprotonation
- Hydrophobic environment → neutral state favored
Implication
- Protein pI cannot be predicted perfectly
- Must be measured experimentally
- Critical for purification (e.g. ion-exchange chromatography)
9. UV Absorption & Protein Concentration 📈

280 nm absorption
- Dominated by tryptophan
- Tyrosine contributes weakly
- Phenylalanine contributes minimally
210–220 nm absorption
- All peptide bonds absorb
- Allows concentration measurement even without aromatics
- Problem: many contaminants also absorb
10. Amino-Acid Analysis (Composition, Not Sequence) 🧪
- Proteins hydrolyzed:
- 110 °C
- 6 M acid
- ~16 h
- Peptide bonds fully broken
- Amino acids derivatized with fluorescent tags
- Separated by chromatography
- Output:
- Which amino acids
- Their relative amounts
- Does not give sequence information
11. Amino-Acid Frequencies in Proteins 📊
- If random: each amino acid ≈ 5%
- Observed deviations:
- Lysine: high frequency → flexible, easy to accommodate
- Cysteine: low frequency → reactive, disulfide risk
- Tryptophan: low frequency → bulky, rigid
12. Levels of Protein Structure 🏗️
- Primary: amino-acid sequence
- Secondary: α-helices, β-sheets, turns
- Tertiary: 3D fold of one chain
- Quaternary: multi-subunit assemblies
13. Peptide Bond Chemistry 🔗
- Formed by:
- Nucleophilic attack of amino group on carboxyl carbon
- Release of water
- Peptide bond has:
- Partial double-bond character
- Planarity
- Dipole moment
- No free rotation around peptide bond
14. Cis vs Trans Peptide Bonds ⚠️
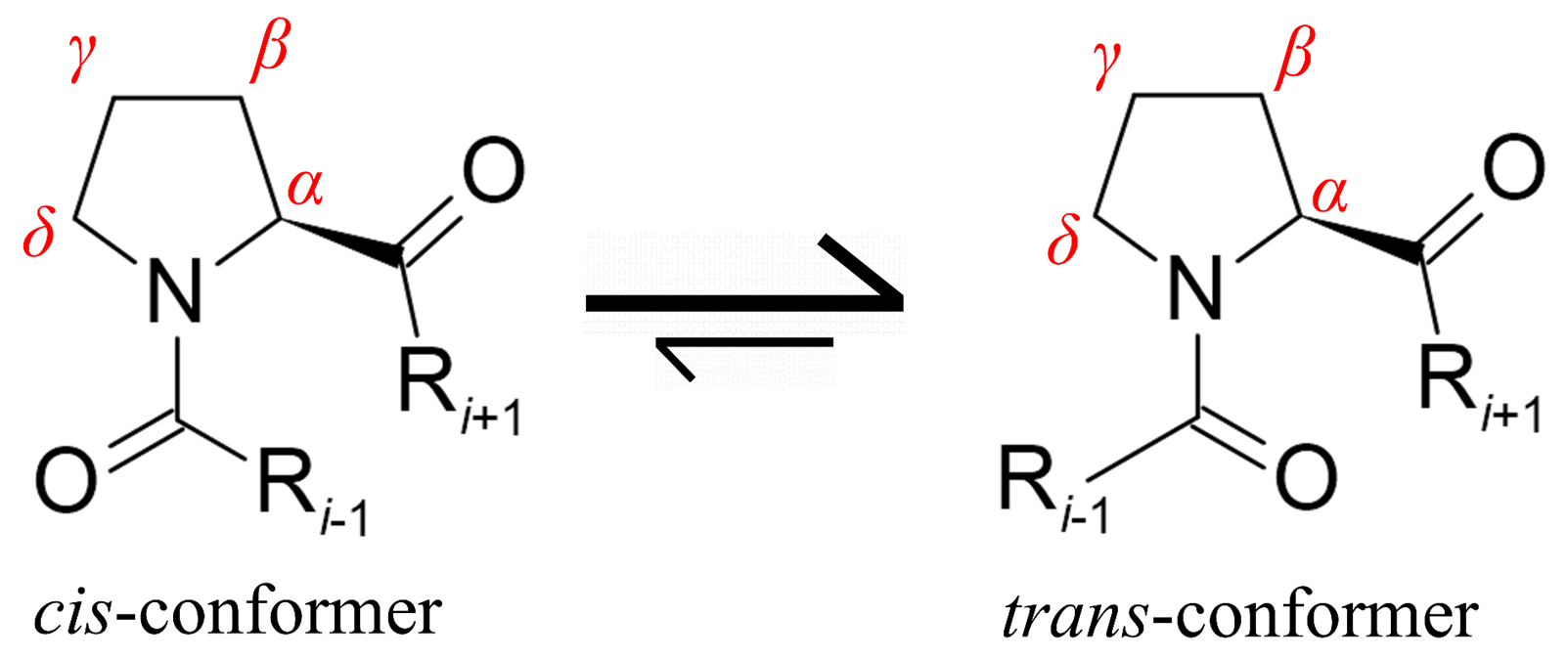

- Trans favored due to sterics
- Exceptions:
- Glycine (tiny)
- Proline (ring locks geometry)
- Cis–trans isomerization of proline:
- Slow
- Requires peptidyl-prolyl isomerases
15. Backbone Angles & Ramachandran Plot 📐


- Two rotatable angles:
- φ (phi): N–Cα
- ψ (psi): Cα–C
- Steric hindrance restricts allowed combinations
- Ramachandran plot shows:
- α-helix regions
- β-sheet regions
- Disallowed zones
16. Secondary Structure Elements 🌀
α-Helix
- 3.6 residues per turn
- Hydrogen bond: i → i+4
- Rise: 1.5 Å per residue
- One turn: 5.4 Å
- Can be amphipathic
- One hydrophobic face
- One hydrophilic face
- Crucial for membrane proteins
β-Sheets
- Parallel or antiparallel
- Hydrogen bonds between strands
- Longer rise per residue (~3.5 Å)
- Connected by turns and loops
17. Amino-Acid Preferences for Secondary Structure 🧩
- α-Helix lovers: Ala, Leu, Met
- β-Sheet lovers: Val, Ile, Phe
- Turn/coil: Gly, Pro
- Proline:
- Breaks helices
- Promotes turns
- Collagen:
- Extremely proline-rich
- Forms specialized triple helices
Final takeaway 🎯
This lecture builds the chemical foundation of proteins:
- Why amino acids behave differently
- How environment controls charge and reactivity
- Why cysteine is rare but powerful
- How structure emerges from chemistry
- Why enzymes position residues precisely
Quiz
Score: 0/30 (0%)



